安装完sightRNA-2.0.0.jar插件
将jar包复制到qupath的extension目录,这个目录是安装后首次启动自定义的。
打开一张需要分析的图:
使用annotation工具圈住要分析的区域:
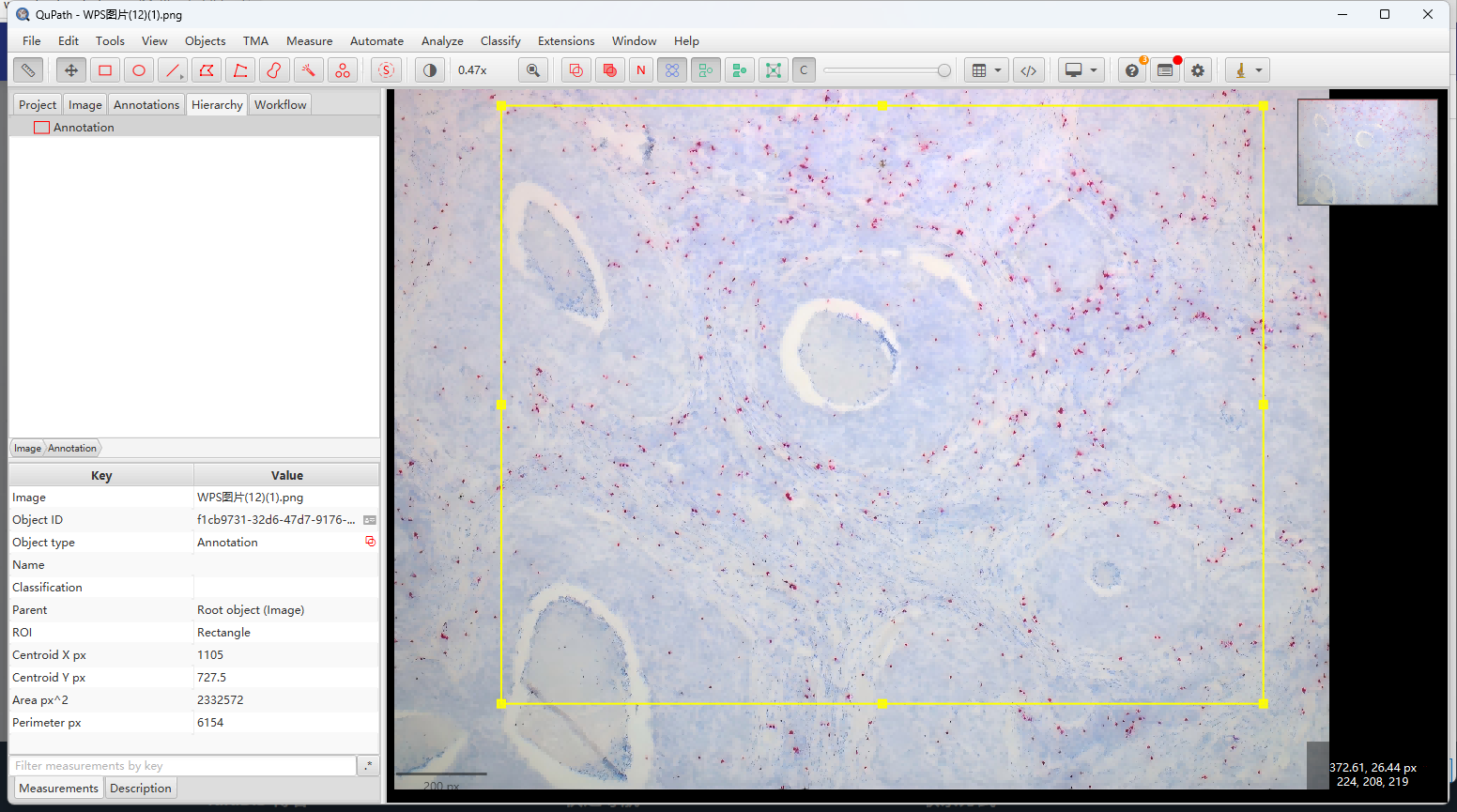
运行Stardist的->Stardist full cell script ,groovy代码如下
/**
* This script provides a general template for cell detection using StarDist in QuPath.
*
* This example assumes you have brightfield image, but want to apply preprocessing
* to separate the stains before running a model trained for fluorescence images.
* One reason to do this is to handle some arbitrary IHC staining if you don't have
* a model trained specifically for your kind of images.
*
* If you use this in published work, please remember to cite *both*:
* - the original StarDist paper (https://doi.org/10.48550/arXiv.1806.03535)
* - the original QuPath paper (https://doi.org/10.1038/s41598-017-17204-5)
*
* There are lots of options to customize the detection - this script shows some
* of the main ones. Check out other scripts and the QuPath docs for more info.
*/
import qupath.ext.stardist.StarDist2D
import qupath.lib.scripting.QP
import qupath.opencv.ops.ImageOps
// IMPORTANT! Replace this with the path to your StarDist model
// that takes a single channel as input (e.g. dsb2018_heavy_augment.pb)
// You can find some at https://github.com/qupath/models
// (Check credit & reuse info before downloading)
def modelPath = "D:/QuPath-0.7.0/model/dsb2018_heavy_augment.pb" //换成你自己的实际地址
// Get current image - assumed to have color deconvolution stains set
var imageData = QP.getCurrentImageData()
var stains = imageData.getColorDeconvolutionStains()
// Customize how the StarDist detection should be applied
// Here some reasonable default options are specified
def stardist = StarDist2D
.builder(modelPath)
.preprocess( // Extra preprocessing steps, applied sequentially (per-tile)
ImageOps.Channels.deconvolve(stains), // Color deconvolution
ImageOps.Channels.extract(0), // Extract the first stain (indexing starts at 0)
ImageOps.Filters.median(2) // Apply a small median filter (optional!)
)
.normalizePercentiles(1, 99) // Percentile normalization
.threshold(0.5) // Probability (detection) threshold
.pixelSize(0.5) // Resolution for detection
.cellExpansion(5) // Expand nuclei to approximate cell boundaries
.measureShape() // Add shape measurements
.measureIntensity() // Add cell measurements (in all compartments)
.build()
// Define which objects will be used as the 'parents' for detection
// Use QP.getAnnotationObjects() if you want to use all annotations, rather than selected objects
def pathObjects = QP.getSelectedObjects()
// Run detection for the selected objects
if (pathObjects.isEmpty()) {
QP.getLogger().error("No parent objects are selected!")
return
}
stardist.detectObjects(imageData, pathObjects)
stardist.close() // This can help clean up & regain memory
println('Done!')
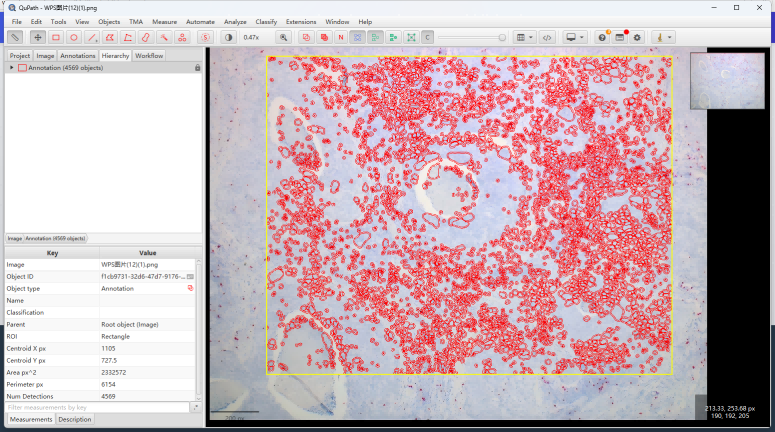
运行完stardist的cell定位,我们开始sightrna的计数分析
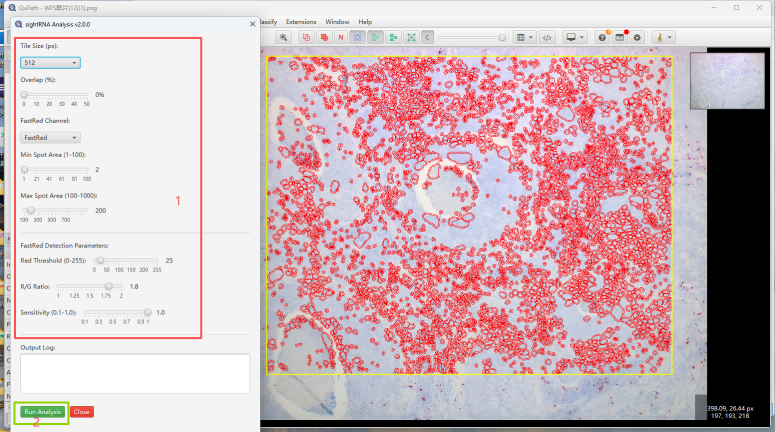
运行的结果文件再用户/sightRNA_results/目录下
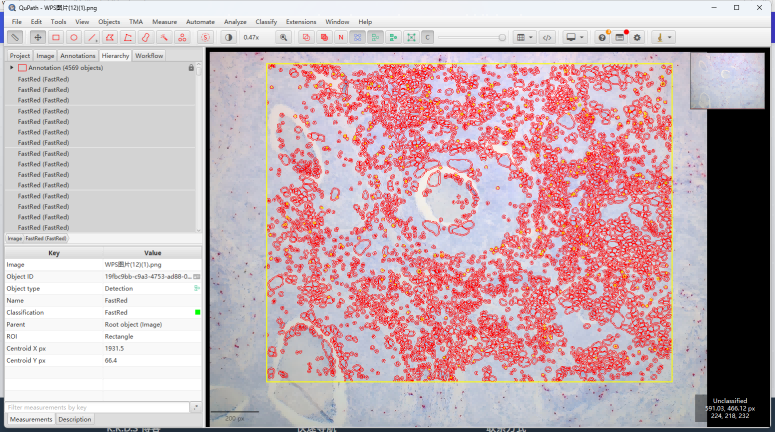
至此完成细胞和fastred的共定位检测。
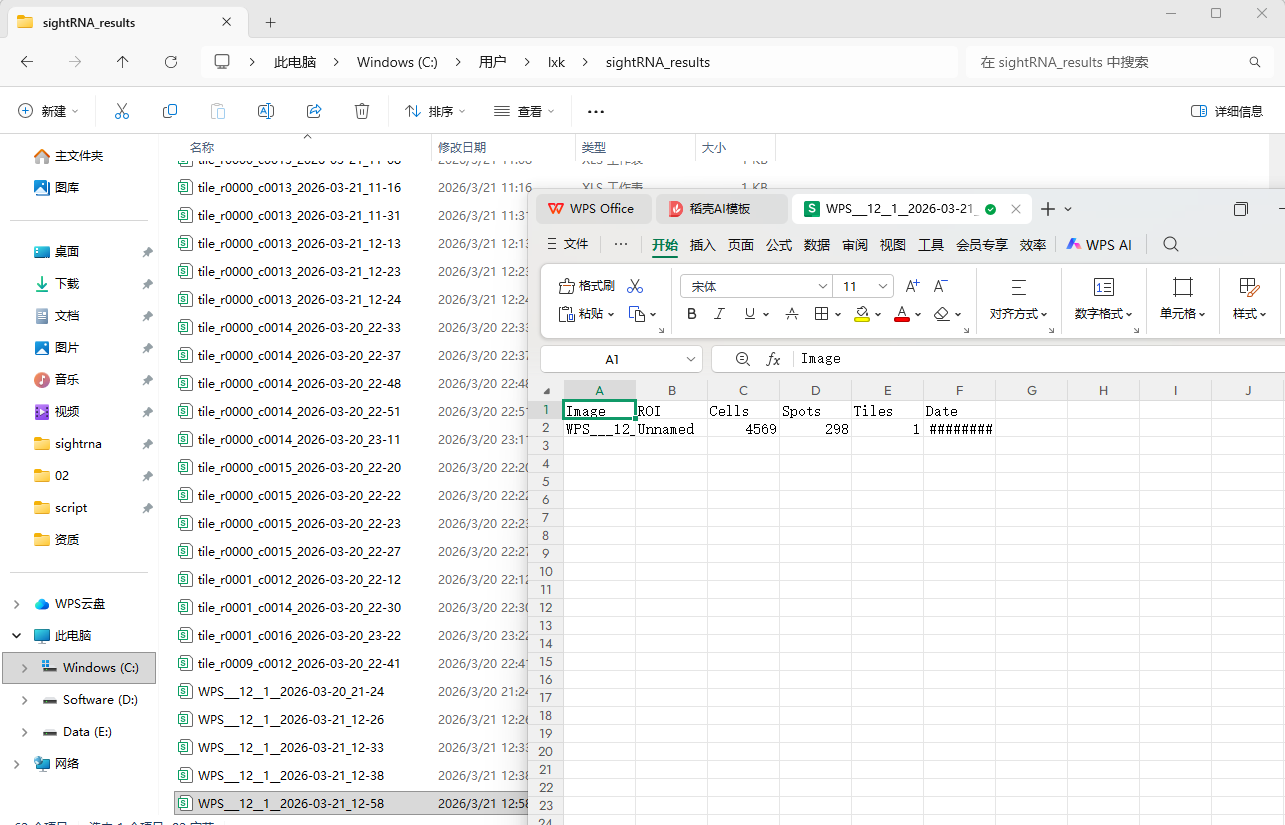
==============================《end》===================================
打完收工!
